Sirtuins, NAD+-dependent signaling and protein (de)acetylation & (de)acylation
Sirtuins, NAD+-dependent signaling and protein (de)acetylation & (de)acylation
Sirtuins (molecule of the month Sept 2017) are an evolutionary conserved enzyme family catalyzing the NAD+-dependent deacylation of protein lysine residues and thereby regulating the functions of these target proteins. The requirement of the central energy metabolite NAD+ as a co-substrate renders Sirtuins metabolic sensors. The seven mammalian Sirtuin isoforms (Sirt1-7) reside in different cellular compartments (Sirt1,6,7: nucleus; Sirt3-5: mitochondria; Sirt2: cytosol) and regulate functions such as energy metabolism, stress responses, and cell proliferation. Sirtuins have been implicated in aging processes and aging-related diseases and are considered attractive targets for the treatment, e.g., of cancer, neurodegenerative disorders, metabolic dysfunctions, and inflammatory diseases. Thus, there is a strong demand for small molecule Sirtuin inhibitors and, in particular, Sirtuin activators for use as chemical tools in functional studies and as lead compounds in drug development.
We study physiological functions and regulation of mammalian Sirtuins as well as their pharmacological modulation with small molecules. Using methods such as activity and binding assays, protein xray crystallography, and mass spectrometry, we identified, e.g., substrate proteins for Sirt1 and Sirt3, the substrate sequence preferences of Sirt1-7, and acyl preferences of Sirt4 and Sirt5. We thereby revealed enzymes involved in energy metabolism and stress responses as Sirt3 targets, rationalizing its roles in processes such as aging-associated hearing loss and metabolic adaptations during exercise; and acylations other than acetylation, such as glutarylation and hydroxymethylglutarylation as preferential substrates for Sirt4 and Sirt5. We further characterized binding sites and modulation mechanisms for a variety of small molecule Sirtuin activators and inhibitors, and identified novel compounds for Sirtuin modulation. We established, e.g., the complex mechanism of Sirtuin inhibition by Ex-527, a compound that entered clinical trials for treatment of Huntington’s Disease, revealing the molecular basis of its specificity. And we contributed to establishing the mechanistic basis of Sirt1 activation and identified first pharmacological activators for Sirt6. The insights in compound mechanisms and the newly identified substances are used by us and others for the rational development of novel, improved Sirtuin modulators.
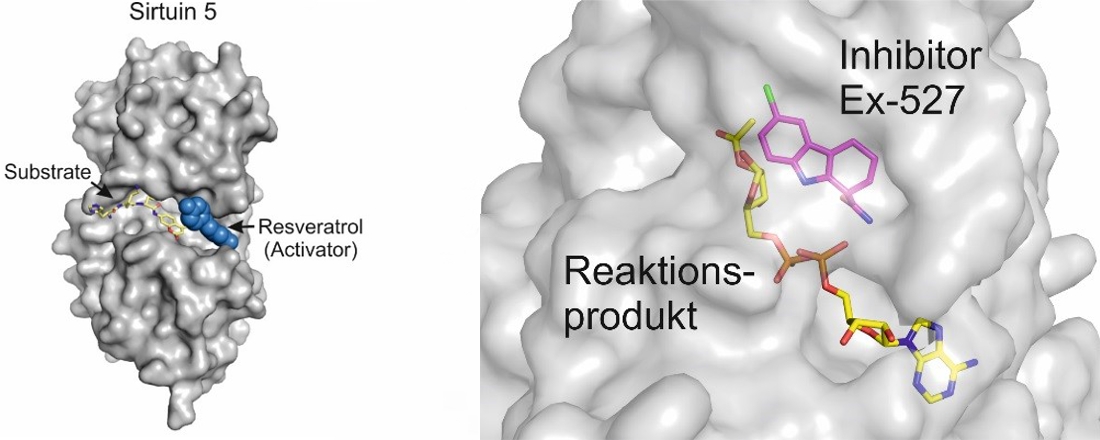
Crystal structures of Sirt5 in complex with a substrate peptide and the activating small molecule resveratrol (left) and of Sirt3 in complex with the rection product 2’-O-acetyl-ADP-ribose and the inhibitor Ex-527.
In a related project, we started to analyze structure, interactions, and mechanisms of Dbc1 (deleted in breast cancer 1), which has been reported to inhibit Sirt1. We found in collaboration with David Sinclair (Harvard Medical School, USA) that Dbc1 also inhibits PARP [poly(adenosine diphosphate–ribose) [PAR] polymerase] 1 (PARP1), another NAD+-dependent signaling enzyme implicated in stress responses. Dbc1 thus co-regulates these two major NAD+-responsive systems. Interestingly, the inhibitory Dbc1/PARP1 interaction can be disrupted by NAD+, presumably through binding to the Dbc1 Nudix domain, establishing the regulation of protein/protein interactions as a novel means to sense NAD+ and Dbc1 as an additional NAD+ sensor in this system. The NAD+-induced disruption of the PARP1/Dbc1 complex activates repair processes and can be stimulated by boosting NAD+ concentrations; a reduction of this effect caused by declining NAD+ levels likely contributes to aging processes.
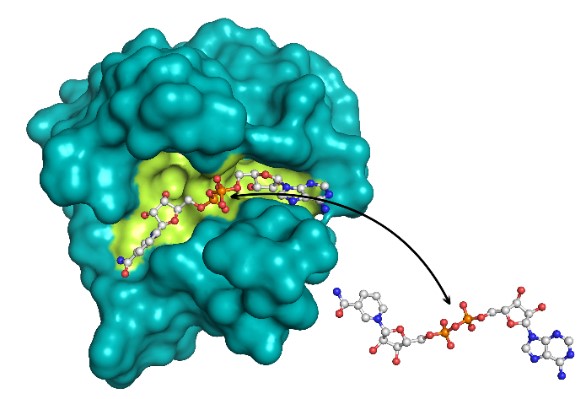
Model of the Dbc1 Nudix domain, with yellow indicating the conserved Nudix dinucleotide binding cleft that appears to mediate binding and sensing of the key energy metabolite NAD+ (ball-and-stick representation)
Selected publications:
M. Pannek, Z. Simic, M. Fuszard, M. Meleshin, D. Rotili, A. Mai, M. Schutkowski, C. Steegborn (2017) Crystal structures of the mitochondrial deacylase Sirtuin 4 reveal isoform-specific acyl recognition and regulation features. Nat. Commun. 8:1513.
J. Li, M.S. Bonkowski, S. Moniot, D. Zhang, B.P. Hubbard, A.J.Y. Ling, L.A. Rajman, B. Qin, Z. Lou, V. Gorbunova, L. Aravind, C. Steegborn, D.A. Sinclair (2017). A conserved NAD+ binding pocket that regulates protein-protein interactions during aging. Science 355, 1312-1317.
W. You, D. Rotili, T.M. Li, C. Kambach, M. Meleshin, M. Schutkowski, K.F. Chua, A. Mai, C. Steegborn (2017) Structural Basis of Sirtuin 6 Activation by Synthetic Small Molecules. Angew. Chem. Int. Ed. Engl. 56, 1007-1011.
D. Rauh, F. Fischer, M. Gertz, M. Lakshminarasimhan, T. Bergbrede, F. Aladini, C. Kambach, C.F.W. Becker, J. Zerweck, M. Schutkowski, C. Steegborn (2013). An acetylome peptide microarray reveals specificities and deacetylation substrates for all human Sirtuin isoforms. Nat. Commun. 4:2327.
M. Gertz, F. Fischer, G.T.T. Nguyen, M. Lakshminarasimhan, M. Schutkowski, M. Weyand, C. Steegborn (2013) Ex-527 inhibits Sirtuins by exploiting their unique NAD+-dependent deacetylation mechanism. Proc. Natl. Acad. Sci. USA 110, E2772-2781.
M. Gertz, G.T. Nguyen, F. Fischer, B. Suenkel, C. Schlicker, B. Fränzel, J. Tomaschewski, F. Aladini, C. Becker, D. Wolters, C. Steegborn (2012) A molecular mechanism for direct sirtuin activation by resveratrol. PLoS One 7:e49761.
